Protein Engineering
We engineer epigenetic proteins through incorporation of unnatural amino acid to capture it’s histone and non-histon binding partners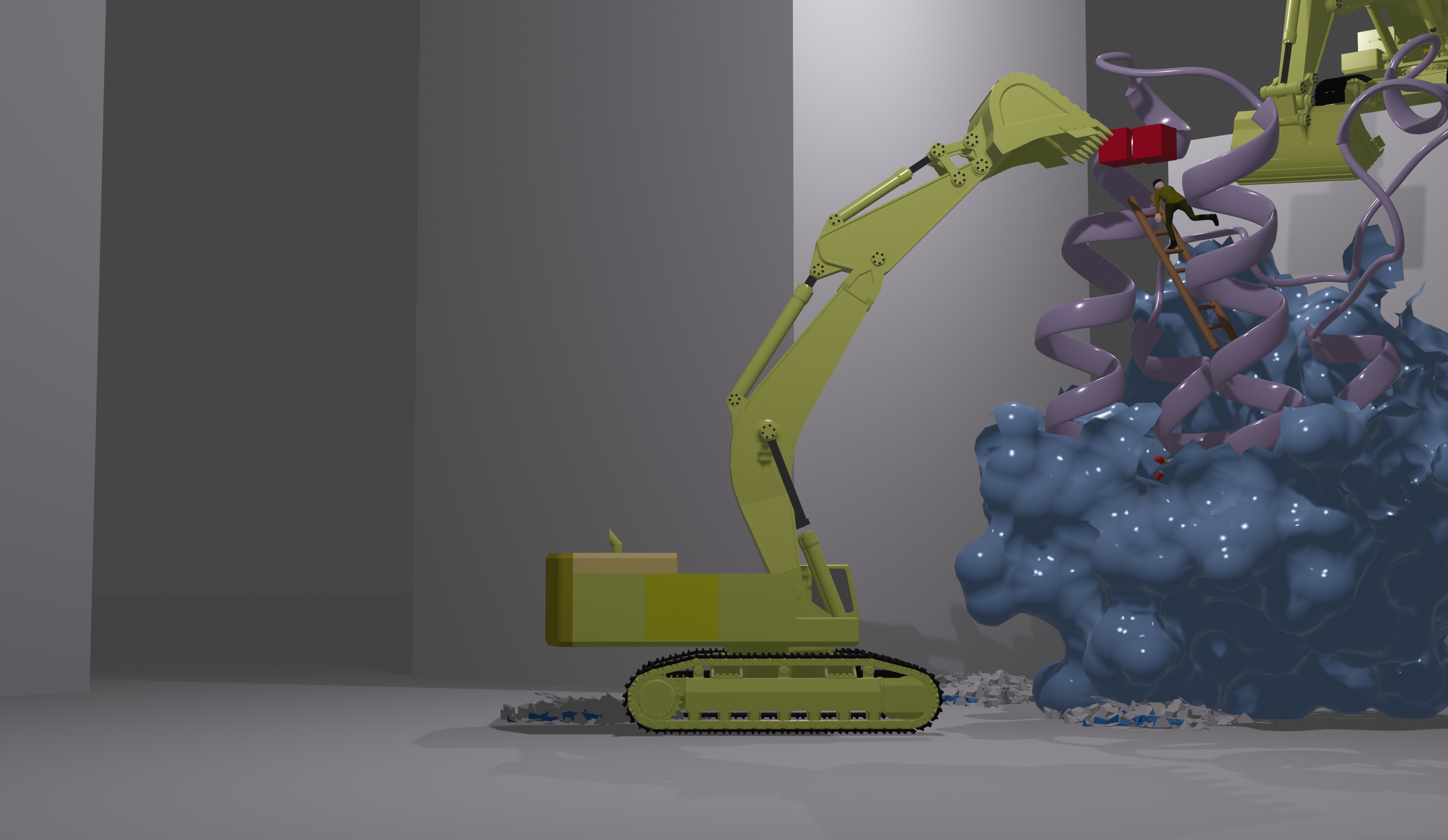 
We engineer epigenetic proteins through incorporation of unnatural amino acid to capture it’s histone and non-histon binding partners 
We have developed a method called as Photo-ChIP which can be used to determine the transient and domain specific protein-DNA interaction 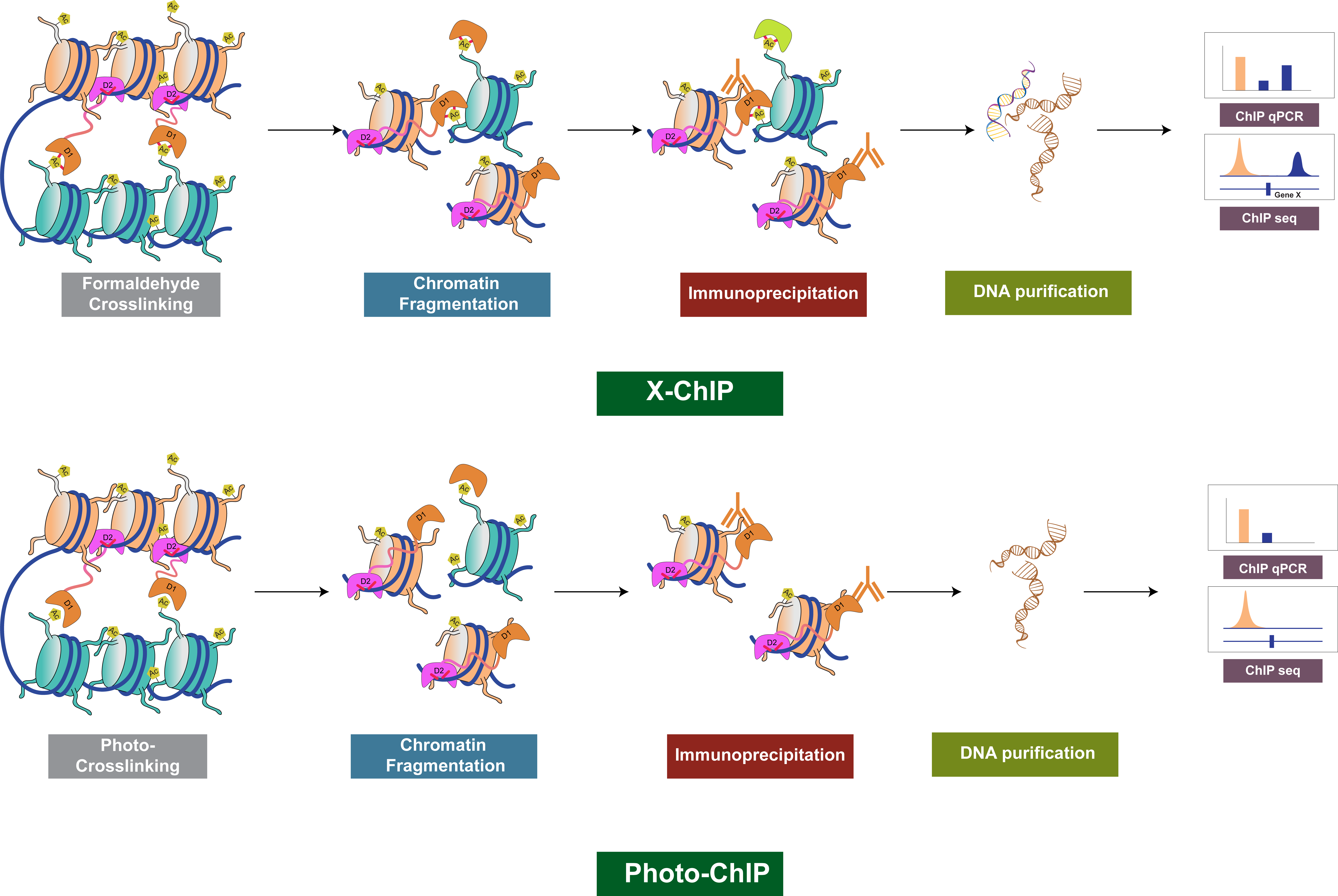
Interaction Based Protein Profiling (IBPP) approach to identify transiently interacting non-histone partners like transcription factors (TFs) and chromatin regulators (CRs). 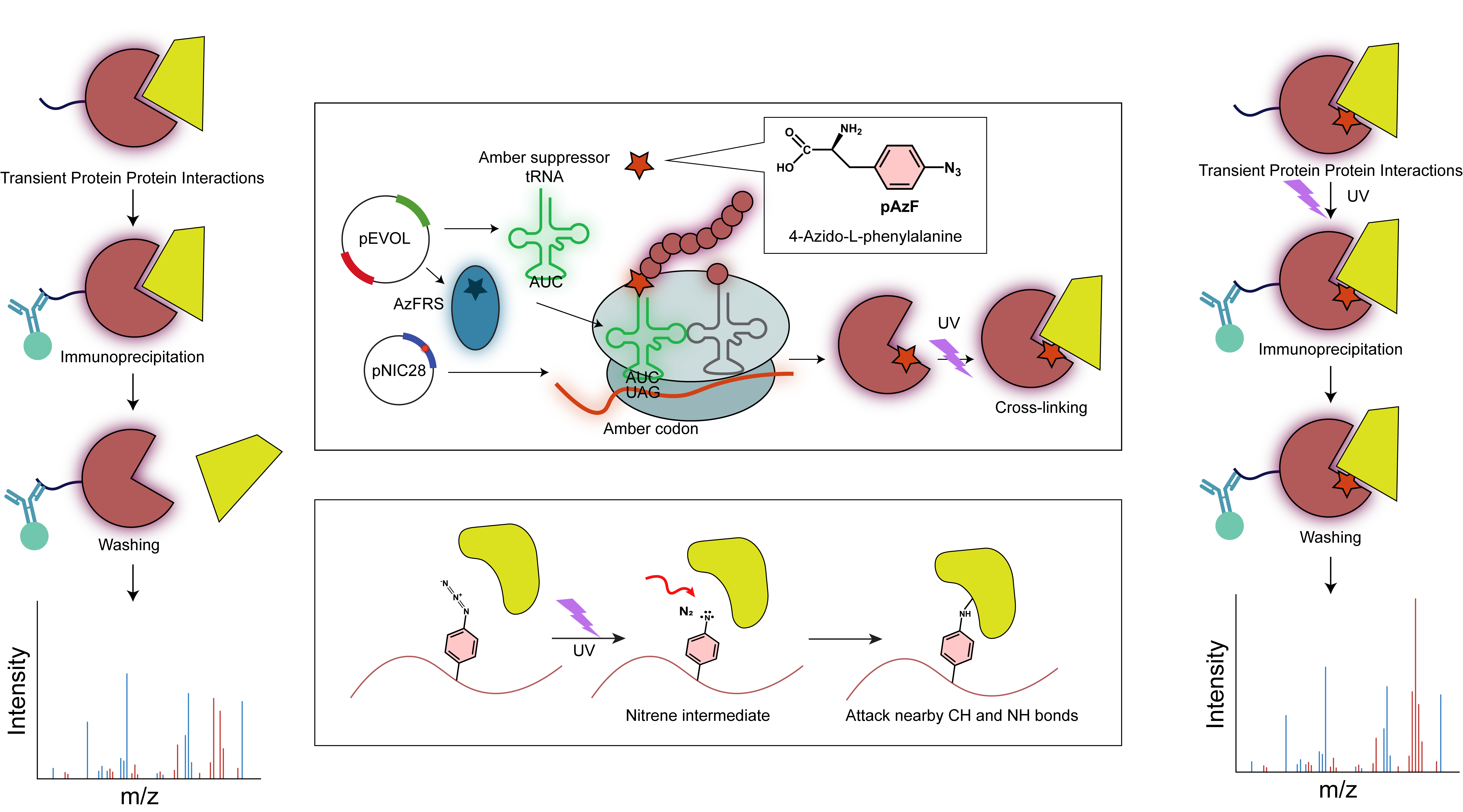
Bromo and Plant Homeodomain Finger Containing Proteins (BRPF) are bromodomain family IV which play as scaffolding proteins and invoves in the regulation of transcription and chromating conformations 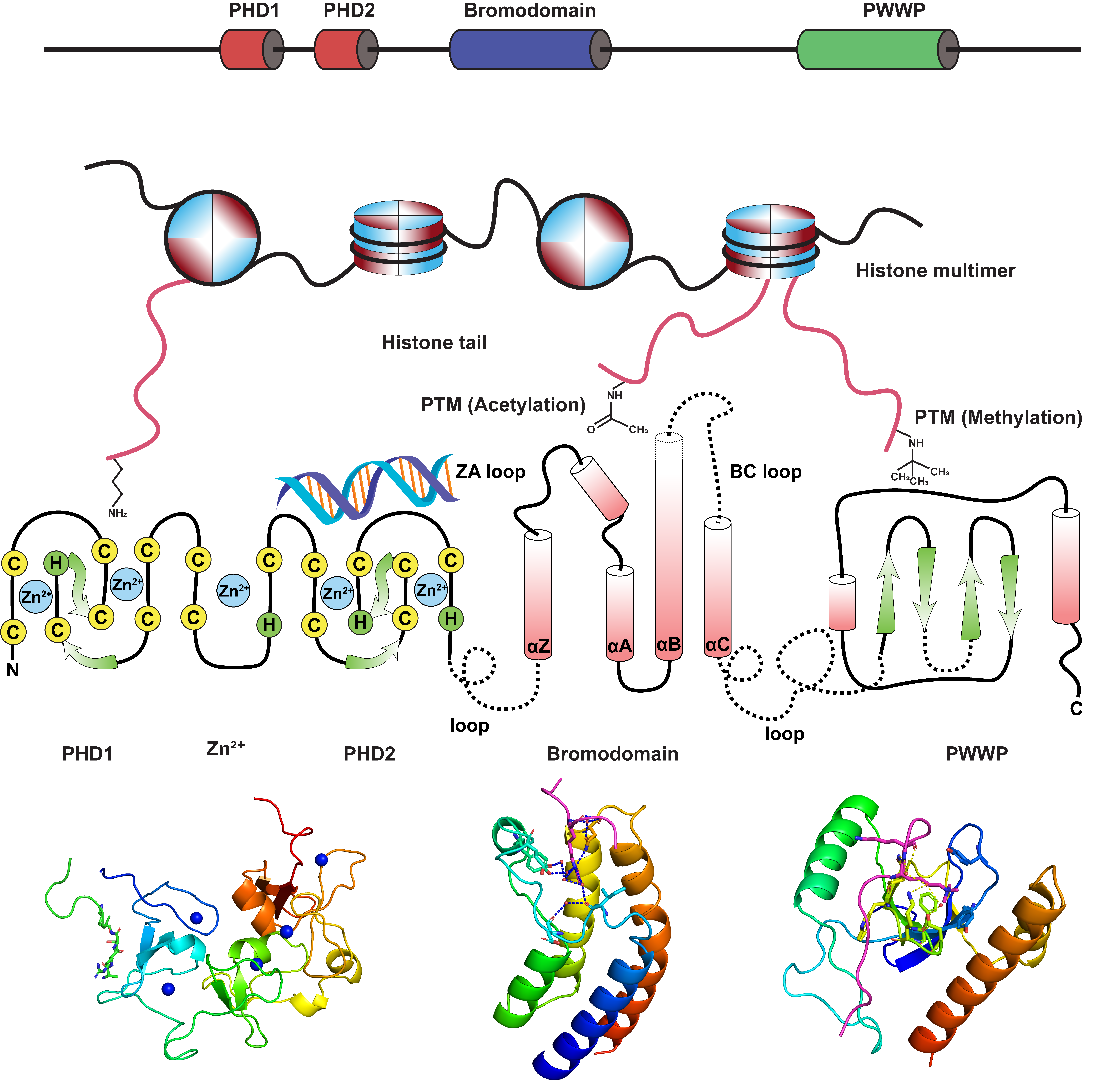
We screen the natural and synthetic small molecule compound libraries to pre-determine potent inhibitors of epigenetic regulators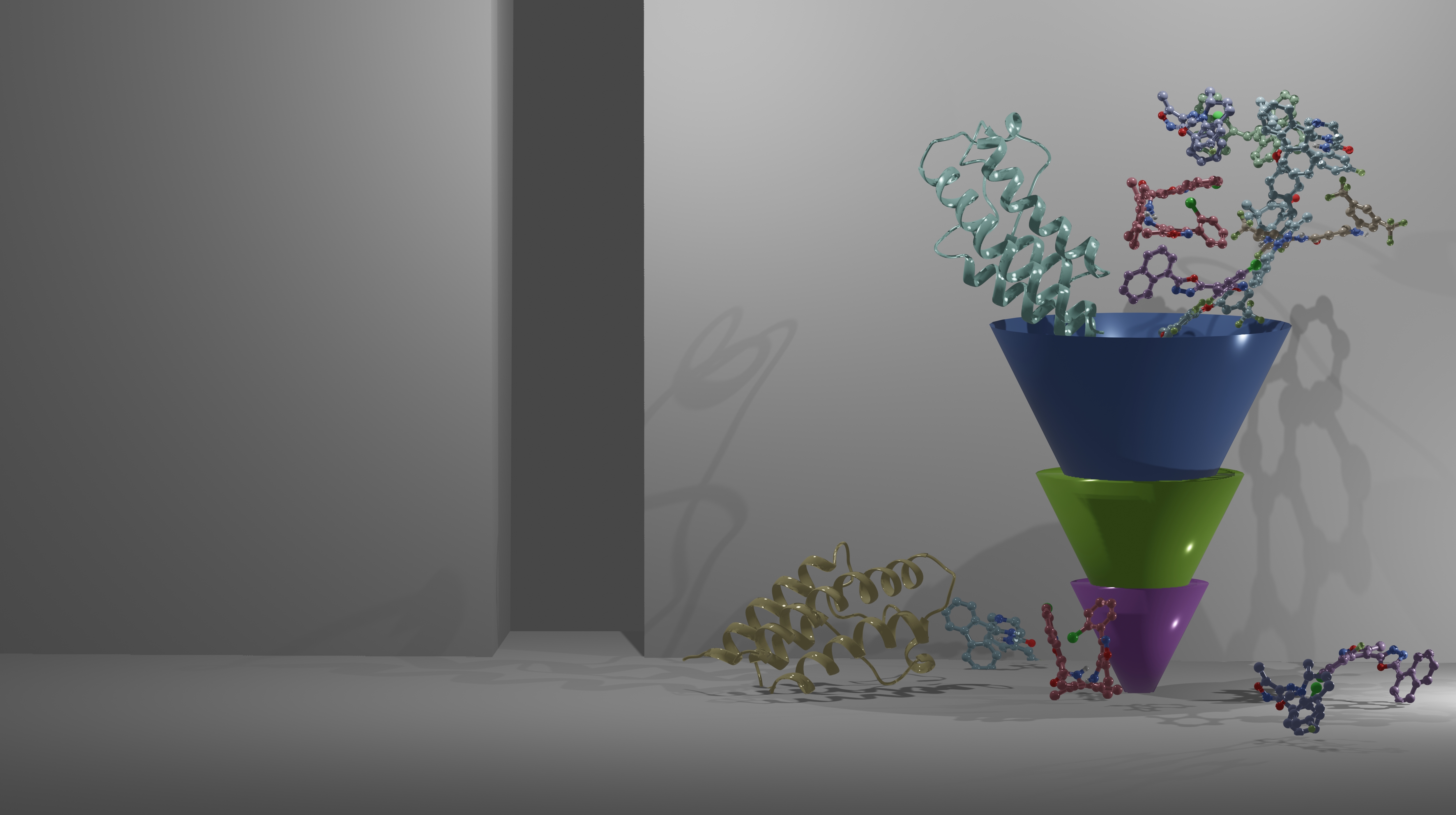 
Published in Chem Commun (Camb), 2021
The site-specific installation of light-activable crosslinker unnatural amino acids offers a powerful approach to trap transient protein–protein interactions both in vitro and in vivo. Herein, we engineer a bromodomain to introduce 4-benzoyl-L-phenylalanine (BzF) using amber suppressor mutagenesis without compromising its ability to recognize the acetylated histone proteins. We demonstrate the high crosslinking efficiency of the engineered reader towards the interacting partners and its suitability for profiling the transient bromodomain interactome.
Recommended citation: Roy A, Barman S, Padhan J, Sudhamalla B. Engineering an acetyllysine reader with a photocrosslinking amino acid for interactome profiling. Chem Commun (Camb). 2021 Sep 28;57(77):9866-9869 https://pubs.rsc.org/en/content/articlelanding/2021/cc/d1cc04611j
Published in Chem Asian J., 2021
BRPF3 is a crucial scaffold protein in gene transcription regulation, forming a complex with HBO1 histone acetyltransferase. Our study reveals that BRPF3 bromodomain interacts with mono- and di-acetylated histone peptides (H4K5ac and H4K5acK12ac). Pull-down assays with full-length histones confirm this interaction. Molecular dynamics simulations illustrate the binding mechanism and stability of the bromodomain-histone peptide complexes. These findings shed light on how BRPF3 directs HBO1 complex recruitment to chromatin for transcriptional regulation.
Recommended citation: Barman S, Roy A, Bardhan I, Kandasamy T, Shivani S, Sudhamalla B. Insights into the Molecular Mechanisms of Histone Code Recognition by the BRPF3 Bromodomain. Chem Asian J. 2021 Nov 2;16(21):3404-3412. https://onlinelibrary.wiley.com/doi/10.1002/asia.202100793
Published in Biochemistry, 2022
BRPF2, a crucial partner of HBO1 histone acetyltransferase complex, plays a role in directing the complex to chromatin for gene expression regulation. Our study unveils BRPF2’s bromodomain as capable of recognizing various acetyllysine residues on histones H4, H3, and H2A, with a preference for H4K5ac, H4K8ac, and H4K5acK12ac modifications. Mutational analysis of the bromodomain identifies key residues for acetyllysine binding. Additionally, ChIP-seq analysis shows BRPF2 co-localizes with HBO1 at H4K5ac and H4K8ac marks near transcription start sites, highlighting its role in chromatin recruitment for gene regulation.
Recommended citation: Barman S, Roy A, Padhan J, Sudhamalla B. Molecular Insights into the Recognition of Acetylated Histone Modifications by the BRPF2 Bromodomain. Biochemistry. 2022 Sep 6;61(17):1774-1789. https://pubs.acs.org/doi/10.1021/acs.biochem.2c00297
Published in J Biomol Struct Dyn, 2022
Bromodomains, vital in gene expression regulation, are implicated in various diseases, making them targets for therapeutic intervention. BRD4, a well-studied bromodomain in cancers like AML and Burkitt Lymphoma, has attracted attention for inhibitor development. However, existing inhibitors often lack specificity and safety. Our study screened natural compounds from the NPASS database against BRD4-BD1 using computational methods. Five inhibitors with comparable or superior binding affinity compared to JQ1 were identified, including NPC268484 (Palodesangren-B), NPC295021 (Candidine), and NPC313112 (Buxifoliadine-D). These findings offer potential candidates for further validation in future experimental studies.
Recommended citation: Barman S, Sahoo SS, Padhan J, Sudhamalla B. Identification of novel natural product inhibitors of BRD4 using high throughput virtual screening and MD simulation. J Biomol Struct Dyn. 2022 Dec 16:1-13. https://www.tandfonline.com/doi/abs/10.1080/07391102.2022.2155346
Published in Biochemistry, 2023
In this study, we investigated how TAF1 dual bromodomain mediates crosstalk among histone modifications. We engineered TAF1 to carry a photoactivatable amino acid, AzF and created analogs targeting each bromodomain separately and both together. Structural integrity was confirmed, and binding properties to acetylated histone ligands were characterized. Both bromodomains independently recognized unique interacting partners at the cellular level. Our findings suggest that BD1 and BD2 of the TAF1 tandem reader independently regulate downstream cellular functions by recognizing specific interacting partners.
Recommended citation: Yadav Y, Barman S#, Roy A#, Padhan J, Sudhamalla B. Uncovering the Domain-Specific Interactome of the TAF1 Tandem Reader Using Site-Specific Azide-Acetyllysine Photochemistry. Biochemistry. 2023 Jan 17;62(2):270-280 https://pubs.acs.org/doi/10.1021/acs.biochem.2c00140
Published in Biomater Sci., 2023
We synthesized block copolymers via RAFT polymerization, containing Boc-protected leucine and Ac-protected glucose. Deprotection yielded amphiphilic polymers forming nano aggregates in water inhibited insulin fibrillation through hydrophobic, electrostatic, and hydrogen bonding interactions. Biophysical techniques confirmed the inhibition, with molecular dynamics simulations supporting the findings.
Recommended citation: Bera A, Ghosh P, Barman S, Bhattacharya S, Sudhamalla B, Goswami K, De P. Insulin fibril inhibition using glycopolymeric nanoassemblies. Biomater Sci. 2023 Mar 31:1-13. https://pubs.rsc.org/en/content/articlelanding/2023/bm/d2bm02078e
Published in Biochem J., 2023
TRIM24 is a chromatin reader that activates estrogen-dependent genes in tumor development. It ubiquitinates p53 and binds a specific histone signature (H3K4me0/H3K23ac) via its RING, PHD, and Bromo domains. High TRIM24 and H3K23ac levels predict poor breast cancer survival. We explored TRIM24 binding to acetylated histone H4 (H4ac). TRIM24 PHD-Bromo preferentially binds H4K5ac, H4K8ac, and H4K5acK8ac. This binding doesn’t interfere with PHD’s recognition of H3K4me0. H4K5ac and H4K8ac co-localize near gene transcription start sites in breast cancer. TRIM24-H4ac targets are linked to vital biological pathways, suggesting a role in transcriptional regulation.
Recommended citation: Bardhan I, Barman S, Roy A, Sudhamalla B. Novel insights into the recognition of acetylated histone H4 tail by the TRIM24 PHD-Bromo module. Biochem J. 2023 May 17; 480 (9): 629-647. https://portlandpress.com/biochemj/article-abstract/480/9/629/232921/Novel-insights-into-the-recognition-of-acetylated?redirectedFrom=fulltext
Published in Journal of Molecular Graphics and Modelling, 2023
Hepatocellular carcinoma (HCC) is a deadly cancer with limited treatment options. We analyzed bromodomain-containing proteins (BRPF) in HCC and found BRPF1 to be significantly upregulated, correlating with poor patient survival. Targeting BRPF1 could improve HCC treatment. We employed high-throughput virtual screening and machine learning-based QSAR modeling to identify new BRPF1 inhibitors. Through extensive molecular dynamics simulations, we discovered four lead compounds with promising binding affinity for the BRPF1 bromodomain, suggesting their potential as potent inhibitors for HCC therapy.
Recommended citation: Barman S, Bardhan I, Padhan J, Sudhamalla B.Integrated virtual screening and MD simulation approaches toward discovering potential inhibitors for targeting BRPF1 bromodomain in hepatocellular carcinoma. Journal of Molecular Graphics and Modelling. 2023, 108642, ISSN 1093-3263. https://www.sciencedirect.com/science/article/pii/S1093326323002401?via%3Dihub
Published in JBC, 2023
BRPF1, a member of the bromodomain-containing protein family, recognizes acetylated histones and non-histone transcription factors to regulate gene expression. While its role in histone recognition is well understood, its regulation via non-histone acetylation remains unexplored. We utilized a protein engineering approach to identify non-histone interactors of BRPF1. By introducing a photo-cross-linkable amino acid into its bromodomain, we efficiently cross-linked interacting partners, revealing novel associations through proteomics. One such partner, interleukin enhancer–binding factor 3, was validated and investigated for its role in hepatocellular carcinoma using ChIP-seq and RNA-seq datasets. These findings enhance our understanding of BRPF1’s transcriptional regulation and aid in developing cancer therapeutics.
Recommended citation: Barman S , Padhan J, Sudhamalla B. Uncovering the Nonhistone Interactome of BRPF1 Bromodomain Using SiteSpecific Azide-Acetyllysine Photochemistry. JBC. 2023 Dec 3; DOI:https://doi.org/10.1016/j.jbc.2023.105551. https://www.sciencedirect.com/science/article/pii/S0021925823025796?via%3Dihub
Published in Bioconjugate Chem., 2024
This study introduces a novel approach to inhibit amyloidosis using peptides derived from the amyloid-promoting sequence of amyloid β-peptide, conjugated with side-chain proline-based methacrylate polymers. These conjugates effectively inhibit lysozyme amyloidosis and reduce cytotoxicity of amyloid aggregations. Synthesized di-, tri-, and tetra-peptide conjugated chain transfer agents (CTAs) were used to polymerize Boc-proline methacryloyloxyethyl ester, resulting in water-soluble polymers with defined peptide chain ends. Among these, the LVFF-conjugated polymer demonstrated potent inhibition of lysozyme amyloidosis, supported by spectroscopic, microscopic, and computational analyses. This study highlights the potential of combining segment-derived amyloid β-peptide sequences with side-chain proline-based polymers for targeting amyloidosis.
Recommended citation: Nayak K, Ghosh P, Barman S, Sudhamalla B, Theato P, De P. Amyloid β-Peptide Segment Conjugated SideChain Proline-Based Polymers as Potent Inhibitors in Lysozyme Amyloidosis. Bioconjugate Chemistry. 2024 Feb 12. https://pubs.acs.org/doi/10.1021/acs.bioconjchem.3c00509
Published:
HBO1 a member of the MYST family of histone acetyltransferases (HATs), was initially identified as a binding partner of ORC that acetylates free histone H3, H4, and nucleosomal H3. It functions as a quaternary complex with the BRPF (BRPF1/2/3) scaffolding protein and two accessory proteins, ING4/5 and Eaf6. Interaction of BRPF2 with HBO1 has been shown to be important for regulating H3K14 acetylation during embryonic development. However, how BRPF2 directs the HBO1 HAT complex to chromatin to regulate its HAT activity toward nucleosomal substrates remains unclear. Our findings reveal novel interacting partners of the BRPF2 bromodomain that recognizes different acetyllysine residues on the N-terminus of histone H4, H3, and H2A and preferentially binds to H4K5ac, H4K8ac, and H4K5acK12ac modifications. In addition, mutational analysis of the BRPF2 bromodomain coupled with isothermal titration calorimetry binding and pull-down assays on the histone substrates identified critical residues responsible for acetyllysine binding. Moreover, the BRPF2 bromodomain could enrich H4K5ac mark-bearing mononucleosomes compared to other acetylated H4 marks. Consistent with this, ChIP-seq analysis revealed that BRPF2 strongly co-localizes with HBO1 at histone H4K5ac and H4K8ac marks near the transcription start sites in the genome. Our study provides novel insights into how the histone binding function of the BRPF2 bromodomain directs the recruitment of the HBO1 HAT complex to chromatin to regulate gene expression.
Published:
HBO1 a member of the MYST family of histone acetyltransferases (HATs), was initially identified as a binding partner of ORC that acetylates free histone H3, H4, and nucleosomal H3. It functions as a quaternary complex with the BRPF (BRPF1/2/3) scaffolding protein and two accessory proteins, ING4/5 and Eaf6. Interaction of BRPF2 with HBO1 has been shown to be important for regulating H3K14 acetylation during embryonic development. However, how BRPF2 directs the HBO1 HAT complex to chromatin to regulate its HAT activity toward nucleosomal substrates remains unclear. Our findings reveal novel interacting partners of the BRPF2 bromodomain that recognizes different acetyllysine residues on the N-terminus of histone H4, H3, and H2A and preferentially binds to H4K5ac, H4K8ac, and H4K5acK12ac modifications. In addition, mutational analysis of the BRPF2 bromodomain coupled with isothermal titration calorimetry binding and pull-down assays on the histone substrates identified critical residues responsible for acetyllysine binding. Moreover, the BRPF2 bromodomain could enrich H4K5ac mark-bearing mononucleosomes compared to other acetylated H4 marks. Consistent with this, ChIP-seq analysis revealed that BRPF2 strongly co-localizes with HBO1 at histone H4K5ac and H4K8ac marks near the transcription start sites in the genome. Our study provides novel insights into how the histone binding function of the BRPF2 bromodomain directs the recruitment of the HBO1 HAT complex to chromatin to regulate gene expression.
Published:
Bromodomain-PHD finger protein 1 (BRPF1) belongs to the BRPF family of bromodomain-containing proteins. Bromodomains are exclusive reader modules that recognize and bind acetylated histones and non-histone transcription factors to regulate gene expression. The biological functions of acetylated histone recognition by BRPF1 bromodomain are well characterized; however, the function of BRPF1 regulation via non-histone acetylation is still unexplored. Therefore, identifying the non-histone interactome of BRPF1 is pivotal in deciphering its role in diverse cellular processes, including its misregulation in diseases like cancer. Herein, we identified the non-histone interacting partners of BRPF1 utilizing a protein engineering-based approach. We site-specifically introduced the unnatural photo-cross-linkable amino acid 4-azido-L-phenylalanine into the bromodomain of BRPF1 without altering its ability to recognize acetylated histone proteins. Upon photoirradiation, the engineered BRPF1 generates a reactive nitrene species, cross-linking interacting partners with spatio-temporal precision. We demonstrated the robust cross-linking efficiency of the engineered variant with reported histone ligands of BRPF1 and further used the variant reader to cross-link its interactome. We also characterized novel interacting partners by proteomics, suggesting roles for BRPF1 in diverse cellular processes. BRPF1 interaction with interleukin enhancer–binding factor 3, one of these novel interacting partners, was further validated by isothermal titration calorimetry and co-IP. Lastly, we used publicly available ChIP-seq and RNA-seq datasets to understand the colocalization of BRPF1 and interleukin enhancer–binding factor 3 in regulating gene expression in the context of hepatocellular carcinoma. Together, these results will be crucial for full understanding of the roles of BRPF1 in transcriptional regulation and in the design of small-molecule inhibitors for cancer treatment.
Undergraduate course, IISER Kolkata, Department of Biological Sciences, 2021
As a Teaching Assistant (TA) for the Cloning and Protein Expression Laboratory, my role encompassed a multifaceted approach to support both the instructor and students in achieving their educational objectives. This laboratory course served as a foundational platform for students to delve into the intricate world of molecular biology techniques, focusing specifically on cloning and protein expression methodologies.
Undergraduate course, IISER Kolkata, Department of Biological Sciences, 2021
As a TA, I provided crucial instructional support during lectures, tutorials, and laboratory sessions, guiding students through the fundamental concepts, algorithms, and tools used in bioinformatics analysis. I facilitated discussions, presented case studies, and conducted hands-on demonstrations to elucidate the application of bioinformatics techniques in biological research.
Undergraduate course, IISER Kolkata, Department of Biological Sciences, 2022
As a Teaching Assistant (TA) for the Cell Biology and Imaging Laboratory, I played a pivotal role in facilitating hands-on learning experiences and fostering a deeper understanding of cellular processes and imaging techniques among students. This laboratory course served as a cornerstone for students to explore the intricate world of cell biology and gain proficiency in microscopy and imaging methodologies.